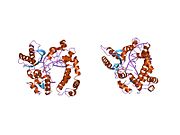DNK polimeraza beta
| DNK polimeraza beta | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
 PDB prikaz baziran na 1bno. | |||||||||||
| Dostupne strukture | |||||||||||
| 1BPX, 1BPY, 1BPZ, 1MQ2, 1MQ3, 1TV9, 1TVA, 1ZJM, 1ZJN, 1ZQA, 1ZQB, 1ZQC, 1ZQD, 1ZQE, 1ZQF, 1ZQG, 1ZQH, 1ZQI, 1ZQJ, 1ZQK, 1ZQL, 1ZQM, 1ZQN, 1ZQO, 1ZQP, 1ZQQ, 1ZQR, 1ZQS, 1ZQT, 2FMP, 2FMQ, 2FMS, 2I9G, 2ISO, 2ISP, 2P66, 2PXI, 3C2K, 3C2L, 3C2M, 3GDX, 3ISB, 3ISC, 3ISD, 3JPN, 3JPO, 3JPP, 3JPQ, 3JPR, 3JPS, 3JPT, 3LK9, 3MBY, 3OGU, 3RH4, 3RH5, 3RH6, 3RJE, 3RJF, 3RJG, 3RJH, 3RJI, 3RJJ, 3RJK, 3TFR, 3TFS, 7ICE, 7ICF, 7ICG, 7ICH, 7ICI, 7ICJ, 7ICK, 7ICL, 7ICM, 7ICN, 7ICO, 7ICP, 7ICQ, 7ICR, 7ICS, 7ICT, 7ICU, 7ICV, 8ICA, 8ICB, 8ICC, 8ICE, 8ICF, 8ICG, 8ICH, 8ICI, 8ICJ, 8ICK, 8ICL, 8ICM, 8ICN, 8ICO, 8ICP, 8ICQ, 8ICR, 8ICS, 8ICT, 8ICU, 8ICV, 8ICW, 8ICX, 8ICY, 8ICZ, 9ICA, 9ICB, 9ICC, 9ICE, 9ICF, 9ICG, 9ICH, 9ICI, 9ICJ, 9ICK, 9ICL, 9ICM, 9ICN, 9ICO, 9ICP, 9ICQ, 9ICR, 9ICS, 9ICT, 9ICU, 9ICV, 9ICW, 9ICX, 9ICY | |||||||||||
| Identifikatori | |||||||||||
| Simboli | POLB; MGC125976 | ||||||||||
| Vanjski ID | OMIM: 174760 MGI: 97740 HomoloGene: 2013 GeneCards: POLB Gene | ||||||||||
| EC broj | 2.7.7.7 | ||||||||||
| |||||||||||
| Pregled RNK izražavanja | |||||||||||
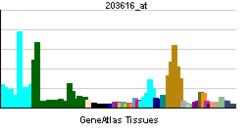 | |||||||||||
| podaci | |||||||||||
| Ortolozi | |||||||||||
| Vrsta | Čovek | Miš | |||||||||
| Entrez | 5423 | 18970 | |||||||||
| Ensembl | ENSG00000070501 | ENSMUSG00000031536 | |||||||||
| UniProt | P06746 | Q8K409 | |||||||||
| RefSeq (mRNA) | NM_002690.2 | NM_011130.2 | |||||||||
| RefSeq (protein) | NP_002681.1 | NP_035260.1 | |||||||||
| Lokacija (UCSC) | Chr 8: 42.2 - 42.23 Mb | Chr 8: 23.74 - 23.76 Mb | |||||||||
| PubMed pretraga | [1] | [2] | |||||||||
| Regulatorni element u POLB | |
|---|---|
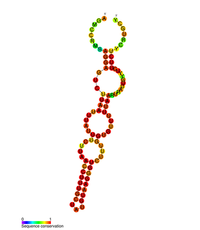 | |
| Predviđena sekundarna struktura stem loopII (M2) regulatornog elementa POLB | |
| Identifikatori | |
| Simbol | POLB |
| Rfam | RF01455 |
| Entrez | 5423 |
| HUGO | POLB |
| OMIM | 174760 |
| RefSeq | NM_002690 |
| Drugi podaci | |
| RNK tip | Cis-reg |
| Domain(i) | Mammalia |
| Lokus | Hromozom 8 p11.2 |
DNK polimeraza beta (POLB) je enzim koji kod ljudi kodira POLB gen.[1]
Funkcija
U eukariotskim ćelijama, DNK polimeraza beta (POLB) izvodi popravke ekscizijom baza (BER) koje su neophodne za popravku DNK, replikaciju, rekombinaciju, i otpornost na lekove.[1]
Regulacija ekspresije
DNK polimeraza beta održava genomski integritet putem uzimanja učešća u popravci ekscizijom baza. Prekomerno izražavanje POLB iRNK je u korelaciji sa brojnim tipovima kancera, dok deficijencije POLB-a rezultuju u hipersenzitivnosti na alkilirajuće agense, uključujući apoptozu, i prekid hromozoma.[2] Iz tih razloga, esencijalno je precizno održavanje kontrole POLB ekspresije.[3][4][5][6]
Interakcije
DNA polimeraza beta formira interakcije sa PNKP[7] i XRCC1.[8][9][10][11]
Reference
- ^ а б „Entrez Gene: POLB polymerase (DNA directed), beta”.
- ^ Narayan S, He F, Wilson SH (1996). „Activation of the human DNA polymerase beta promoter by a DNA-alkylating agent through induced phosphorylation of cAMP response element-binding protein-1”. J. Biol. Chem. 271 (31): 18508—13. PMID 8702497. doi:10.1074/jbc.271.31.18508.
- ^ Y, Canitrot; C, Cazaux; Fréchet M; et al. (1998). „Overexpression of DNA polymerase beta in cell results in a mutator phenotype and a decreased sensitivity to anticancer drugs”. Proc. Natl. Acad. Sci. U.S.A. 95 (21): 12586—90. PMC 22874
 . PMID 9770529. doi:10.1073/pnas.95.21.12586.
. PMID 9770529. doi:10.1073/pnas.95.21.12586. - ^ V, Bergoglio; MJ, Pillaire; Lacroix-Triki M; et al. (2002). „Deregulated DNA polymerase beta induces chromosome instability and tumorigenesis”. Cancer Res. 62 (12): 3511—4. PMID 12067997.
- ^ V, Bergoglio; Y, Canitrot; Hogarth L; et al. (2001). „Enhanced expression and activity of DNA polymerase beta in human ovarian tumor cells: impact on sensitivity towards antitumor agents”. Oncogene. 20 (43): 6181—7. PMID 11593426. doi:10.1038/sj.onc.1204743.
- ^ Srivastava DK, Husain I, Arteaga CL, Wilson SH (1999). „DNA polymerase beta expression differences in selected human tumors and cell lines”. Carcinogenesis. 20 (6): 1049—54. PMID 10357787. doi:10.1093/carcin/20.6.1049.
- ^ Whitehouse, C J; Taylor R M; Thistlethwaite A; Zhang H; Karimi-Busheri F; Lasko D D, Weinfeld M; Caldecott K W (2001). „XRCC1 stimulates human polynucleotide kinase activity at damaged DNA termini and accelerates DNA single-strand break repair”. Cell. United States. 104 (1): 107—17. ISSN 0092-8674. PMID 11163244. doi:10.1016/S0092-8674(01)00195-7.
- ^ Wang, Liming; Bhattacharyya Nandan; Chelsea Diane M; Escobar Pedro F, Banerjee Sipra (2004). „A novel nuclear protein, MGC5306 interacts with DNA polymerase beta and has a potential role in cellular phenotype”. Cancer Res. United States. 64 (21): 7673—7. ISSN 0008-5472. PMID 15520167. doi:10.1158/0008-5472.CAN-04-2801.
- ^ Fan, Jinshui; Otterlei Marit (2004). „XRCC1 co-localizes and physically interacts with PCNA”. Nucleic Acids Res. England. 32 (7): 2193—201. PMC 407833
 . PMID 15107487. doi:10.1093/nar/gkh556.
. PMID 15107487. doi:10.1093/nar/gkh556. - ^ Kubota, Y; Nash R A (1996). „Reconstitution of DNA base excision-repair with purified human proteins: interaction between DNA polymerase beta and the XRCC1 protein”. EMBO J. ENGLAND. 15 (23): 6662—70. ISSN 0261-4189. PMC 452490
 . PMID 8978692.
. PMID 8978692. - ^ Bhattacharyya, N; Banerjee S (2001). „A novel role of XRCC1 in the functions of a DNA polymerase beta variant”. Biochemistry. United States. 40 (30): 9005—13. ISSN 0006-2960. PMID 11467963. doi:10.1021/bi0028789.
Literatura
- T, Date; Tanihara K; Yamamoto S; et al. (1992). „Two regions in human DNA polymerase beta mRNA suppress translation in Escherichia coli”. Nucleic Acids Res. 20 (18): 4859—64. PMC 334243
 . PMID 1408801. doi:10.1093/nar/20.18.4859.
. PMID 1408801. doi:10.1093/nar/20.18.4859. - Wang L, Patel U, Ghosh L, Banerjee S (1992). „DNA polymerase beta mutations in human colorectal cancer”. Cancer Res. 52 (17): 4824—7. PMID 1511447.
- T, Tokui; Inagaki M; Nishizawa K; et al. (1991). „Inactivation of DNA polymerase beta by in vitro phosphorylation with protein kinase C”. J. Biol. Chem. 266 (17): 10820—4. PMID 2040602.
- DN, SenGupta; Zmudzka BZ; Kumar P; et al. (1986). „Sequence of human DNA polymerase beta mRNA obtained through cDNA cloning”. Biochem. Biophys. Res. Commun. 136 (1): 341—7. PMID 2423078. doi:10.1016/0006-291X(86)90916-2.
- Zmudzka BZ, Fornace A, Collins J, Wilson SH (1988). „Characterization of DNA polymerase beta mRNA: cell-cycle and growth response in cultured human cells”. Nucleic Acids Res. 16 (20): 9587—96. PMC 338765
 . PMID 2460824. doi:10.1093/nar/16.20.9587.
. PMID 2460824. doi:10.1093/nar/16.20.9587. - Widen SG, Kedar P, Wilson SH (1988). „Human beta-polymerase gene. Structure of the 5'-flanking region and active promoter”. J. Biol. Chem. 263 (32): 16992—8. PMID 3182828.
- J, Abbotts; SenGupta DN; Zmudzka B; et al. (1988). „Expression of human DNA polymerase beta in Escherichia coli and characterization of the recombinant enzyme”. Biochemistry. 27 (3): 901—9. PMID 3284575. doi:10.1021/bi00403a010.
- Y, Dobashi; Kubota Y; Shuin T; et al. (1995). „Polymorphisms in the human DNA polymerase beta gene”. Hum. Genet. 95 (4): 389—90. PMID 7705833.
- YJ, Chyan; Ackerman S; Shepherd NS; et al. (1994). „The human DNA polymerase beta gene structure. Evidence of alternative splicing in gene expression”. Nucleic Acids Res. 22 (14): 2719—25. PMC 308239
 . PMID 7914364. doi:10.1093/nar/22.14.2719.
. PMID 7914364. doi:10.1093/nar/22.14.2719. - Maruyama K, Sugano S (1994). „Oligo-capping: a simple method to replace the cap structure of eukaryotic mRNAs with oligoribonucleotides”. Gene. 138 (1–2): 171—4. PMID 8125298. doi:10.1016/0378-1119(94)90802-8.
- M, Chang; Burmer GC; Sweasy J; et al. (1994). „Evidence against DNA polymerase beta as a candidate gene for Werner syndrome”. Hum. Genet. 93 (5): 507—12. PMID 8168825.
- Chyan YJ, Strauss PR, Wood TG, Wilson SH (1996). „Identification of novel mRNA isoforms for human DNA polymerase beta”. DNA Cell Biol. 15 (8): 653—9. PMID 8769567. doi:10.1089/dna.1996.15.653.
- H, Pelletier; Sawaya MR; Wolfle W; et al. (1996). „Crystal structures of human DNA polymerase beta complexed with DNA: implications for catalytic mechanism, processivity, and fidelity”. Biochemistry. 35 (39): 12742—61. PMID 8841118. doi:10.1021/bi952955d.
- H, Pelletier; Sawaya MR; Wolfle W; et al. (1996). „A structural basis for metal ion mutagenicity and nucleotide selectivity in human DNA polymerase beta”. Biochemistry. 35 (39): 12762—77. PMID 8841119. doi:10.1021/bi9529566.
- Pelletier H, Sawaya MR (1996). „Characterization of the metal ion binding helix-hairpin-helix motifs in human DNA polymerase beta by X-ray structural analysis”. Biochemistry. 35 (39): 12778—87. PMID 8841120. doi:10.1021/bi960790i.
- Y, Kubota; Nash RA; Klungland A; et al. (1997). „Reconstitution of DNA base excision-repair with purified human proteins: interaction between DNA polymerase beta and the XRCC1 protein”. EMBO J. 15 (23): 6662—70. PMC 452490
 . PMID 8978692.
. PMID 8978692. - Bennett RA, Wilson DM, Wong D, Demple B (1997). „Interaction of human apurinic endonuclease and DNA polymerase beta in the base excision repair pathway”. Proc. Natl. Acad. Sci. U.S.A. 94 (14): 7166—9. PMC 23779
 . PMID 9207062. doi:10.1073/pnas.94.14.7166.
. PMID 9207062. doi:10.1073/pnas.94.14.7166. - MR, Sawaya; Prasad R; Wilson SH; et al. (1997). „Crystal structures of human DNA polymerase beta complexed with gapped and nicked DNA: evidence for an induced fit mechanism”. Biochemistry. 36 (37): 11205—15. PMID 9287163. doi:10.1021/bi9703812.
- Bhattacharyya N, Banerjee S (1997). „A variant of DNA polymerase beta acts as a dominant negative mutant”. Proc. Natl. Acad. Sci. U.S.A. 94 (19): 10324—9. PMC 23361
 . PMID 9294209. doi:10.1073/pnas.94.19.10324.
. PMID 9294209. doi:10.1073/pnas.94.19.10324. - Y, Suzuki; Yoshitomo-Nakagawa K; Maruyama K; et al. (1997). „Construction and characterization of a full length-enriched and a 5'-end-enriched cDNA library”. Gene. 200 (1–2): 149—56. PMID 9373149. doi:10.1016/S0378-1119(97)00411-3.
Vidi još
- POLA1
- POLA2
Spoljašnje veze
- п
- р
- у
-
 1bno: NMR struktura N-terminalnog domena DNK polimeraze beta
1bno: NMR struktura N-terminalnog domena DNK polimeraze beta -
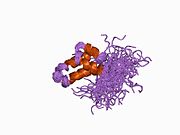 1bnp: NMR struktura N-terminalnog domena DNK polimeraze beta, 55 struktura
1bnp: NMR struktura N-terminalnog domena DNK polimeraze beta, 55 struktura -
 1bpb: Kristalna struktura DNK polimeraze beta pacova
1bpb: Kristalna struktura DNK polimeraze beta pacova -
 1bpd: Kristalna struktura DNK polimeraze beta pacova
1bpd: Kristalna struktura DNK polimeraze beta pacova -
 1bpe: Kristalna struktura DNK polimeraze beta pacova
1bpe: Kristalna struktura DNK polimeraze beta pacova -
 1bpx: Kompleks DNK polimeraze beta i DNK
1bpx: Kompleks DNK polimeraze beta i DNK -
 1bpy: Ljudska DNK polimeraza beta u kompleksu sa oštećenom DNK i DDCTP
1bpy: Ljudska DNK polimeraza beta u kompleksu sa oštećenom DNK i DDCTP -
 1bpz: Ljudska DNK polimeraza beta u kompleksu sa oštećenom DNK
1bpz: Ljudska DNK polimeraza beta u kompleksu sa oštećenom DNK -
 1dk2: Struktura N-terminalnog domena DNK polimeraze beta
1dk2: Struktura N-terminalnog domena DNK polimeraze beta -
 1dk3: Struktura N-terminalnog domena DNK polimeraze beta
1dk3: Struktura N-terminalnog domena DNK polimeraze beta -
 1huo: Kristalna struktura DNK polimeraze beta u kompleksu sa DNK i CR-TMPPCP
1huo: Kristalna struktura DNK polimeraze beta u kompleksu sa DNK i CR-TMPPCP -
 1huz: Kristalna struktura DNK polimeraze beta u kompleksu sa DNK i CR-PCP
1huz: Kristalna struktura DNK polimeraze beta u kompleksu sa DNK i CR-PCP